Deep Learning for Image Segmentation of Tomato Plants
目次
Motivation
At Incubit we are always looking for new ways to apply AI and machine learning technology to interesting problems. A recent project has found us applying image segmentation to the problem of identifying parts of a tomato plant. Tomate pruning is a technique often used to increase the yield of tomato plants by removing small, non-tomato-blooming branches from the plant. Here we describe a method to determine the location of prunable tomato branches, as well as critical parts of the tomato plant which should not be touched, such as primary trunks, branches supporting tomatoes, and the tomatoes themselves.

Figure 1: Pruning non-critical branches from a tomato plant. Photo courtesy of gardeningknowhow.com.
In this project we apply techniques for Image Segmentation to locate objects of interest. The goal of this analysis method is to locate different segments, or contiguous sets of pixels, within an image which denote some meaningful entity within the image. There are various ways to accomplish this using both computer vision and model-based approaches. We opted for the supervised model-based approach, with the hopes of obtaining both higher accuracy and greater generalization to new images.
Obtaining labeled data
The first step in developing an image model is obtaining labeled training data. For this task we used Incubit’s AI Platform (IAP) to create segmentation labels. Figure 2 shows an example of how we annotated an image to show segments of four classes of interest: main trunk, sucker branch, tomato branch, and tomato.

Figure 2: On left – raw image. On right – annotated image. Annotations are: red = main trunk, blue = sucker branch, purple = tomato branch, yellow = tomato.
These annotations were stored as JSON files and used to create segmentation masks. Figure 3 shows an example of these masks, created from a crop of the above annotations, which was fed directly into the model as labels:

Figure 3: Masks created from the annotated data. The white pixels in each mask act as the target for the segmentation of each respective class.
The model
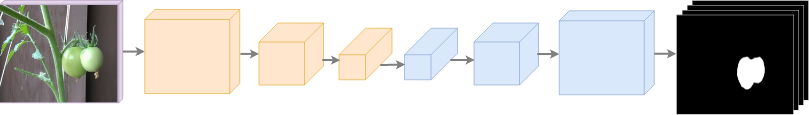
We based on model architecture on SegNet, a well-known deep neural network which excels in image segmentation applications. Figure 4 shows a simplified overview of the architecture used.

A high-level summary of the architecture is this: the original image is passed through a number of encoding blocks, each of which consists of several convolutional layers, batch normalization, and ReLU activation layers, followed by a pooling layer. The reduced features are then passed through a series of upsampling layers. Loss is computed from the cross entropy between the sigmoid output of the final convolutional layer and the segmentation targets (labels).
Training
Training was performed with a constant learning rate of 0.00001 until there was no improvement in the test error rate for 10 consecutive epochs. A random parameter search yielded the following combination of optimal hyperparameters: 5 encoding and decoding blocks, 32 initial filters, a dropout rate of 0.25, and independent pooling indexes between the encoding and decoding layers.
Image augmentation was used to both increase the size of the training pool and to help generalize the model. A combination of flips, crops, random noise, Gaussian blur, fine and coarse dropout, perspective transformations, piecewise affines, rotations, shears, and elastic transformations were used from the imgaug library to reach this end.
Figure 5 shows an example of an annotated output frame produced by the trained model.

Figure 5: Annotated output showing different segmentation classes of a tomato plant.
Post-Processing
One of the expected outcomes of this project is the ability to automatically locate the origin and direction of branches growing from the main trunk. To do this, we can utilize the segmentation outputs for the trunk and branch classes. We wrote an algorithm to detect branches which are attached to the main trunk and to extract this information. The steps are:
- Use the connected components algorithm to identify individual branches and trunks within the image.
- For each possible pair of branch and trunk segments which partially overlap (full overlaps represent non-connected branch/trunk pairs), record these as connected pairs.
- For each connected pair, mark the base of the branch as the centroid of the overlapped region. This acts as the starting point of the branch direction vector.
- Define the end point of the branch direction vector as the halfway point of the shortest line connecting the following entities:
- The least-squares fit line of the branch segment
- The centroid of the branch segment
Figure 6 shows an example of a branch direction vector drawn from the base of the branch, at the trunk, to the midpoint of the branch:

Figure 6: Drawing a branch direction vector.
Branch vectors, along with the segmented classes, are superimposed on the original raw image.
Result
Here is a video showing the results of this analysis on a tomato garden. Visible are the different class segments and the branch direction vectors.
We look forward to applying this technology to other interesting and novel use cases.


